SHRIMP (Scalable Heuristic Robust Iterative Matrix Profile) is a heuristic algorithm for computing the Matrix Profile with large scale time series in very short time. The project paper can be read here
Install depdencies using Poetry.
# install poetry package manager
pip3 install poetry
# install project dependencies
poetry installPerformance comparison results can be summarized using command line interface in src/main.py:
cd src/
python3 main.py -n 100000 -m 5000 --scrump --shrimpThis should output the following plots:
This should output the following plots incidating the results of each algorithm, and how it compares to the true matrix profile (as calculated by STUMP), as well as statistics of each algorthim, runtime (in seconds), Mean-Squared-Error, Median-Squared-Error, and Convergence Percentage:
usage: main.py [-h] [-n N] [-m M] [--shrimp] [--scrump] [--stamp] [--scrimp] [--shrimp_hill] [--shrimp_mass] [--shrimp_diag] [--shrimp_ga]
options:
-h, --help show this help message and exit
-n N length of time series
-m M window size
--shrimp run SHRIMP
--scrump run SCRUMP
--stamp run STAMP
--scrimp run SCRIMP
--shrimp_hill run SHRIMP_HILL
--shrimp_mass run SHRIMP_MASS
--shrimp_diag run SHRIMP_DIAG
--shrimp_ga run SHRIMP_GA
See Algorithms for more information on how to further tune each of these algorithms and their hyperparameters.
To run all tests:
pytest -v src/tests.pyIn order to configure individual algorithms, edit the algorithms variable in src/main.py based on the configuration information found below for the corresponding algorithm.
algorithms = [
...
{
'name': 'SHRIMP',
'callable': SHRIMP,
'kwargs': {
'step_size': int(m / 2),
'max_aspirations': 2 * int(np.sqrt(n - m + 1)),
'mass_percentage': 1.0,
},
},
...
]STUMP is the JIT-compiled, parallelized Matrix Profile computation module from library stumpy, implementing the STOMPopt algorithm with Pearson correlations. STUMP is always executed in the src/main.py as a baseline measure for the ground-truth.
SHRIMP is the JIT-compiled, parallelized Matrix Profile computation, introduced in this project, and evolved from SHRIMP_DIAG.
{
'name': 'SHRIMP',
'callable': SHRIMP,
'kwargs': {
'step_size': int(m / 2), # initial step size for first-pass compuation of MASS
'max_aspirations': 2 * int(np.sqrt(n - m + 1)), # size of aspriations list, i.e. number of "best distances" to store
'mass_percentage': 1.0, # fraction of vertical column to include in MASS computation
},
},SCRUMP is the JIT-compiled, parallelized Matrix Profile computation module from library stumpy, implementing SCRIMP/SCRIMP++.
{
'name': 'SCRUMP',
'callable': scrump_callable,
'kwargs': {
'iterations': 5, # number of times to update the matrix profile by computing new distances (calls update() method)
},
},STAMP is the Matrix Profile computation algorithm, defined in this paper.
{
'name': 'STAMP',
'callable': STAMP,
'kwargs': {
'completion': 0.5, # fraction of random columns of the distance matrix to be computed
},
},STOMP is the Matrix Profile computation algorithm, defined in this paper.
{
'name': 'STOMP',
'callable': STOMP,
'kwargs': {}, # STOMP provides no tunable hyperparameters
},SCRIMP is the Matrix Profile computation algorithm, defined in this paper.
{
'name': 'SCRIMP',
'callable': SCRIMP,
'kwargs': {
'completion': 0.075, # fraction of diagonals of the distance_matrix to be computed
},
},SHRIMP_MASS is most-promising-minimum column search algorithm for computing Matrix Profile introduced in this project.
{
'name': 'SHRIMP_MASS',
'callable': SHRIMP_MASS,
'kwargs': {
'steps': m, # initial step size for first-pass compuation of MASS
'max_iter': 100, # maximum iterations for computing "current-best" distances
'search_window': m // 4, # search window size for computing MASS around "current-best" distance
},
},SHRIMP_HILL is most-promising-minimum hill climbing search algorithm for computing Matrix Profile introduced in this project.
{
'name': 'SHRIMP_HILL',
'callable': SHRIMP_HILL,
'kwargs': {
'steps': m, # initial step size for first-pass compuation of MASS
'max_iter': 175, # maximum iterations for computing "current-best" distances
'epsilon': 0.003, # amount of change in z-normalized-euclidian-distance before the "hill-climbing" deems it insignifiant in improvement
},
},SHRIMP_DIAG is most-promising-minimum diagonal hill climbing search algorithm for computing Matrix Profile introduced in this project.
{
'name': 'SHRIMP_DIAG',
'callable': SHRIMP_DIAG,
'kwargs': {
'step_size': m // 2, # initial step size for first-pass compuation of MASS
'max_aspirations': 5 * int(math.sqrt(n - m + 1)), # size of aspriations list, how many current-best" distances we should compute diagonals for
},
},SHRIMP_DIAG is genetic algorithm for computing Matrix Profile introduced in this project.
{
'name': 'SHRIMP_GA',
'callable': SHRIMP_GA,
'kwargs': {
'pop_size': 500, # number of agents in the population
'generations': 200, # number of generations to simulation
'mutate': 0.05 # random mutation chance
},
},- The first half of
src/algos.pycontains the implementations for the STAMP, STOMP, and SCRIMP++ algorthims as described in their respective papers. The second half ofsrc/algos.pycontains the preliminary implementations our SHRIMP algorithm, including methods that compute the matrix profile using MASS based off of the current lowest distance (SHRIMP_MASS), as well as hill climbing (SHRIMP_HILL), and finally a diagonal-based computation method based on hill-climbing (SHRIMP_DIAG, which is the basis for our final algorithm). src/utils.pycontains helper functions that are critical to the computation of the matrix profile, including the aforementioned MASS function, and z-normalized-euclidian-distance functions.src/shrimp.pycontains the current final version of our SHRIMP algorithm, using JIT, and multi-threading.src/genetic.pycontains a preliminary implementation of our SHRIMP algorithm using Genetic Algorithms as a basis for search. It performed below expectations and was dropped from testing.src/preliminary_results.pyis a precursor to thesrc/final_results.pyscript, acting as our first test-bench during preliminary testing.src/final_results.pyis the script used to generate the final results and comparisons.src/hm.pyis a spin-off to the main testbench as instead of generating a graph showing best distance vectors, it instead shows a heatmap of all distances between the two time-series, as well as which indicides are computed by the algorithm.
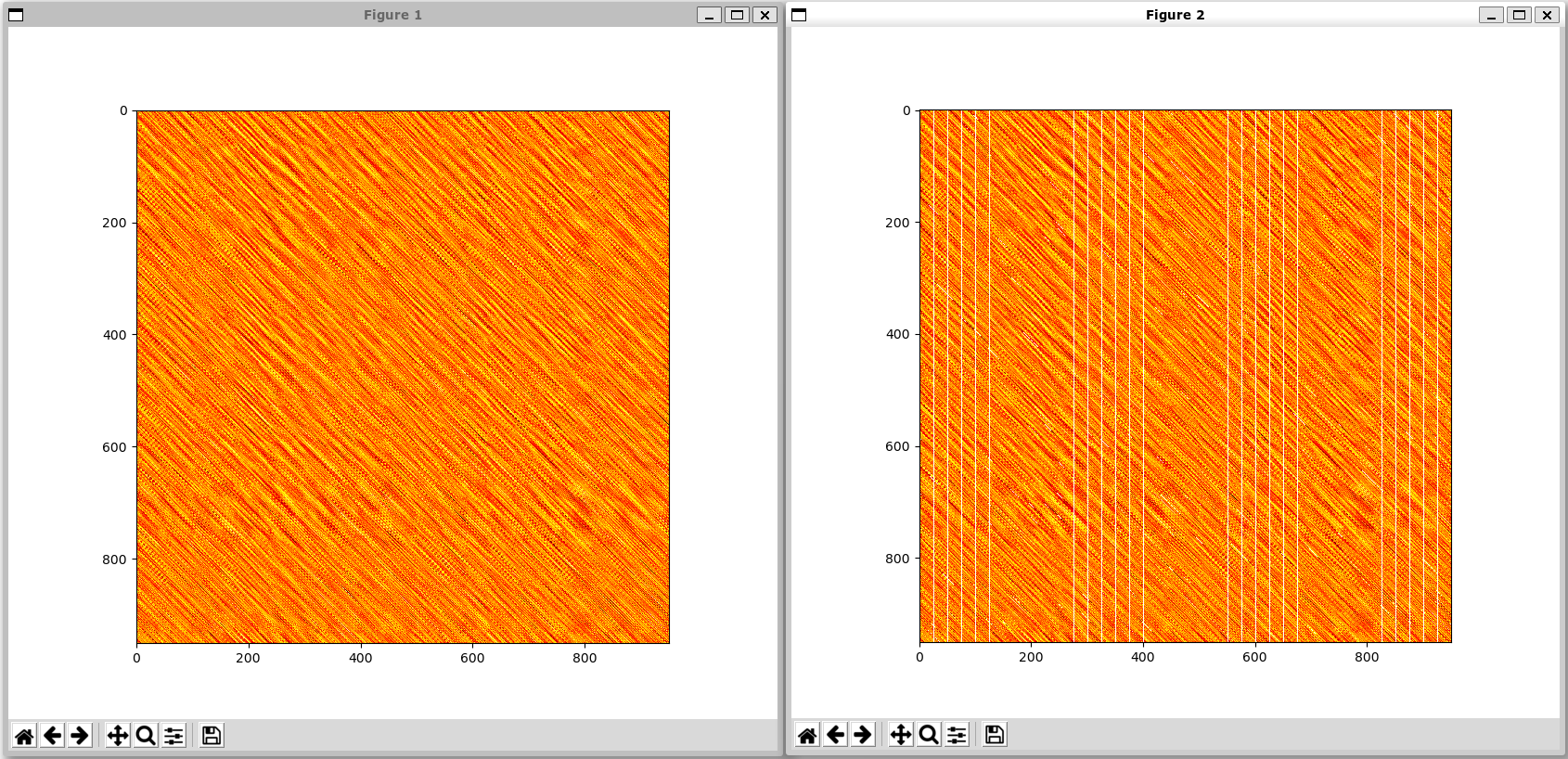
Figure 1 on the left shows the distance matrix of all pairs of incidies between the two time-series, and Figure 2 on the right shows which indicies the algorithm has done computations on. Note that only SHRIMP_MASS, SHRIMP_DIAG, and SHRIMP_GA support generating the visited heatmaps.